Articles /Network Graph Analysis /Part 5
LLM-Augmented Knowledge Graphs
Using large language models to extract structured knowledge and build navigable graphs
Knowledge GraphsLLMPythonNetworkX
Articles /Network Graph Analysis /Part 5
Knowledge GraphsLLMPythonNetworkX
Large language models excel at extracting structured information from text -- turning paragraphs into entities, relations, and hierarchies that map directly to graph structures. This article explores how LLMs serve as knowledge extractors, how prompt design shapes the quality of extracted triples, and how LLM outputs become navigable graphs. The approach builds on the Books project ↗'s technique of converting LLM-generated outlines into NetworkX trees.
Traditional NLP pipelines chain NER, coreference resolution, and relation extraction -- each introducing error that compounds downstream. An LLM can perform end-to-end extraction in a single prompt: given a passage, output a list of (subject, relation, object) triples. The quality depends on prompt design, but for many domains LLM extraction matches or exceeds pipeline approaches with far less engineering.
The extraction prompt needs to specify the entity types of interest, the relation vocabulary (open or closed), and the output format. A closed schema (fixed entity types and relation labels) produces cleaner graphs at the cost of missing novel relationships. An open schema captures more but requires post-processing to merge duplicates and normalize relation names. Few-shot examples in the prompt improve consistency significantly -- showing the model 3-5 input/output pairs anchors its behavior more reliably than long instructions alone.
Beyond flat triples, LLMs can generate hierarchical outlines -- a book's structure as nested topics, a codebase as module trees, a curriculum as prerequisite chains. The Books project ↗ demonstrates this: an LLM summarizes a book into a structured outline, which is then parsed into a tree graph. The key is constraining the output format (JSON, indented text, or markdown headings) so the parser can reliably convert text to graph.
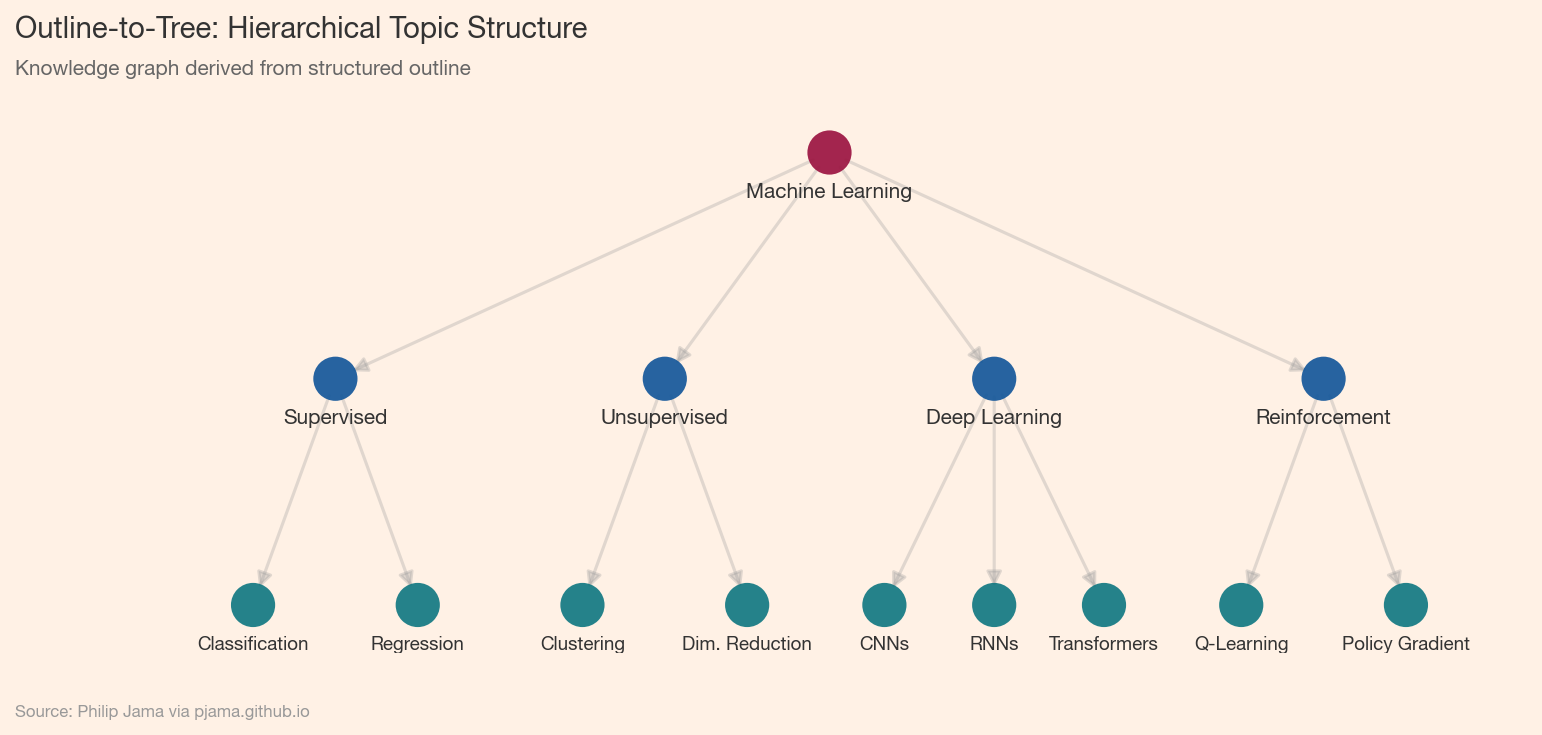
Parsing LLM output into graphs requires handling indentation levels (for nested lists), section numbering (for headings), or JSON structure (for explicit hierarchies). Each indent level or nesting depth maps to a parent-child edge. Robustness comes from normalizing whitespace, handling edge cases (empty sections, inconsistent formatting), and validating the resulting graph (connected, acyclic for trees).
LLM-extracted graphs are only as good as the extraction prompt and the post-processing pipeline. Common failure modes include hallucinated entities (the model invents nodes not present in the source), missed relations (especially implicit ones), and inconsistent naming (the same entity appears under multiple surface forms). Entity resolution -- merging "United States", "US", and "America" into a single node -- is essential for producing clean graphs. Embedding-based deduplication works well here: embed all extracted entity names and merge pairs above a cosine similarity threshold.
The next article extends these LLM-built knowledge graphs into a retrieval system: GraphRAG replaces traditional vector search with graph traversal, using the relational structure to retrieve richer, more connected context for generation.
These LLM-built knowledge graphs encode rich structure -- but how do you query them at generation time? Graph-based retrieval replaces vector search with traversal.
If you're exploring related work and need hands-on help, I'm open to consulting and advisory. Get in touch›