Articles /Network Graph Analysis /Part 4
Building Knowledge Graphs from Text
From raw text to structured concept networks
Knowledge GraphsNLPPythonNetwork Analysis
Articles /Network Graph Analysis /Part 4
Knowledge GraphsNLPPythonNetwork Analysis
Text is full of implicit structure: entities, relationships, hierarchies that a knowledge graph makes explicit. By extracting concepts and their connections from documents, we transform unstructured prose into a navigable network of ideas. This article covers the pipeline from text to graph, drawing on the concept extraction and associative network techniques used in the Graphception project ↗.
A knowledge graph represents information as a network of entities (nodes) connected by labeled relationships (edges). Unlike a flat database table, a knowledge graph captures the structure of knowledge: how concepts relate, which ideas are central, and where clusters of related topics form. Examples range from Wikidata and Google’s Knowledge Graph to domain-specific ontologies in medicine, law, and engineering.
The first step is pulling structured triples (subject, relation, object) from text. Approaches range from rule-based (dependency parsing + patterns) to statistical (named entity recognition + relation classification) to neural (end-to-end transformer models). The choice depends on domain, corpus size, and required precision.
The simplest knowledge graph is a co-occurrence network: two concepts share an edge if they appear together (in a sentence, paragraph, or document). This captures topical association but not the type of relationship. A semantic graph adds labeled, directed edges (e.g., causes, part-of, treats) that encode meaning. Co-occurrence networks are easy to build; semantic graphs require deeper NLP but yield richer reasoning.
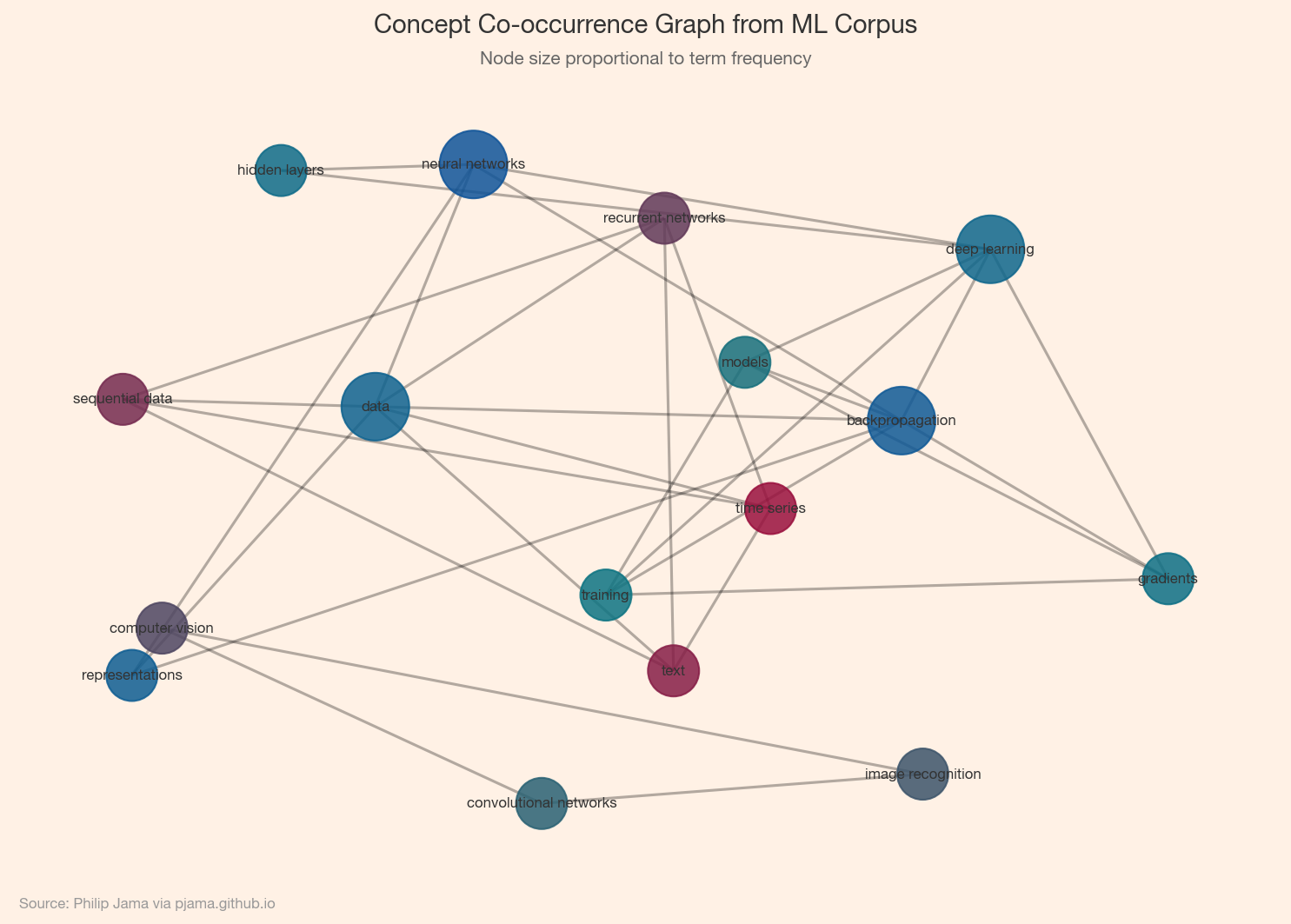
Once you have a concept graph, community detection (Part 2) reveals topic clusters: groups of concepts that co-occur frequently. These clusters often correspond to subtopics or themes within the corpus. Visualizing them helps identify the main threads in a body of text and the bridging concepts that connect different domains. The Graphception project ↗ demonstrates this pipeline on real text corpora.
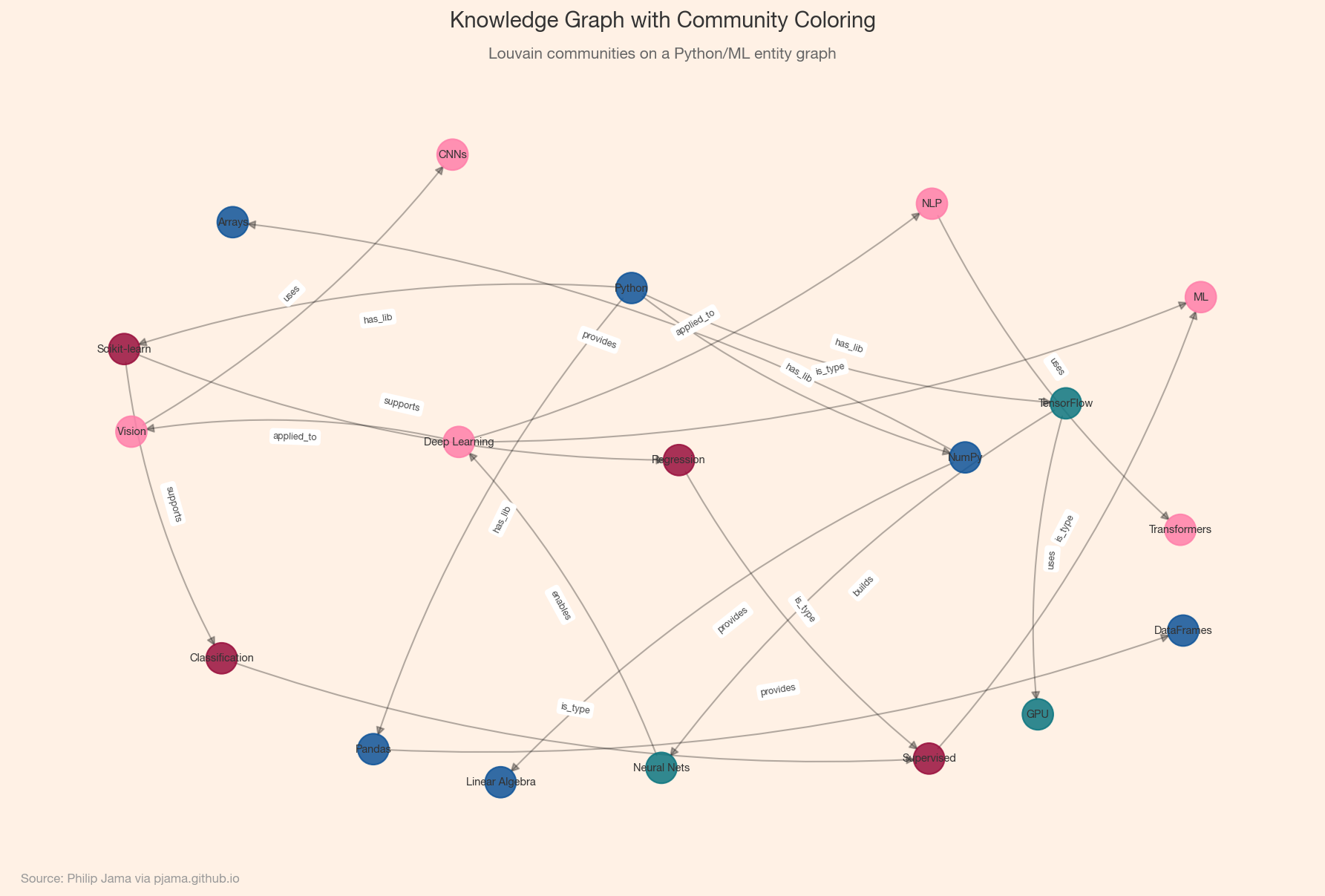
Small knowledge graphs fit in memory as NetworkX objects. Larger graphs benefit from graph databases (Neo4j, Amazon Neptune) or RDF triple stores (Apache Jena). For analysis at scale, adjacency-list formats (edge lists, CSR matrices) and distributed frameworks (GraphX, DGL) keep things tractable. The choice of storage shapes what queries are efficient: traversals, pattern matching, or bulk analytics.
Co-occurrence and NLP pipelines produce structured graphs, but they miss implicit relations. Large language models can fill those gaps.
If you're exploring related work and need hands-on help, I'm open to consulting and advisory. Get in touch›