Articles /Network Graph Analysis /Part 8
Temporal Graph Networks
Modeling graphs that evolve over time
Temporal GraphsGraph Neural NetworksDeep LearningPython
Articles /Network Graph Analysis /Part 8
Temporal GraphsGraph Neural NetworksDeep LearningPython
Real networks change: friendships form and dissolve, citations accumulate, communication patterns shift with the calendar. Static graph analysis captures a snapshot, but temporal graph networks model the dynamics: how structure and features evolve, and how past interactions inform future ones. Building on the GNN foundations from Part 7 (Graph Neural Networks), this article introduces temporal graph representations, dynamic random walks, and the architectures (TGAT, TGN) that learn from time-stamped interactions.
A static graph is a single photograph; a temporal graph is a film. Edges appear and disappear, node attributes change, and the patterns that matter are often sequential: a burst of activity, a gradual drift, a sudden rewiring. Capturing these dynamics requires representations that encode when things happen, not just what is connected.
Two main representations:
(u, v, t, features). More expressive but requires specialized architectures that process events incrementally.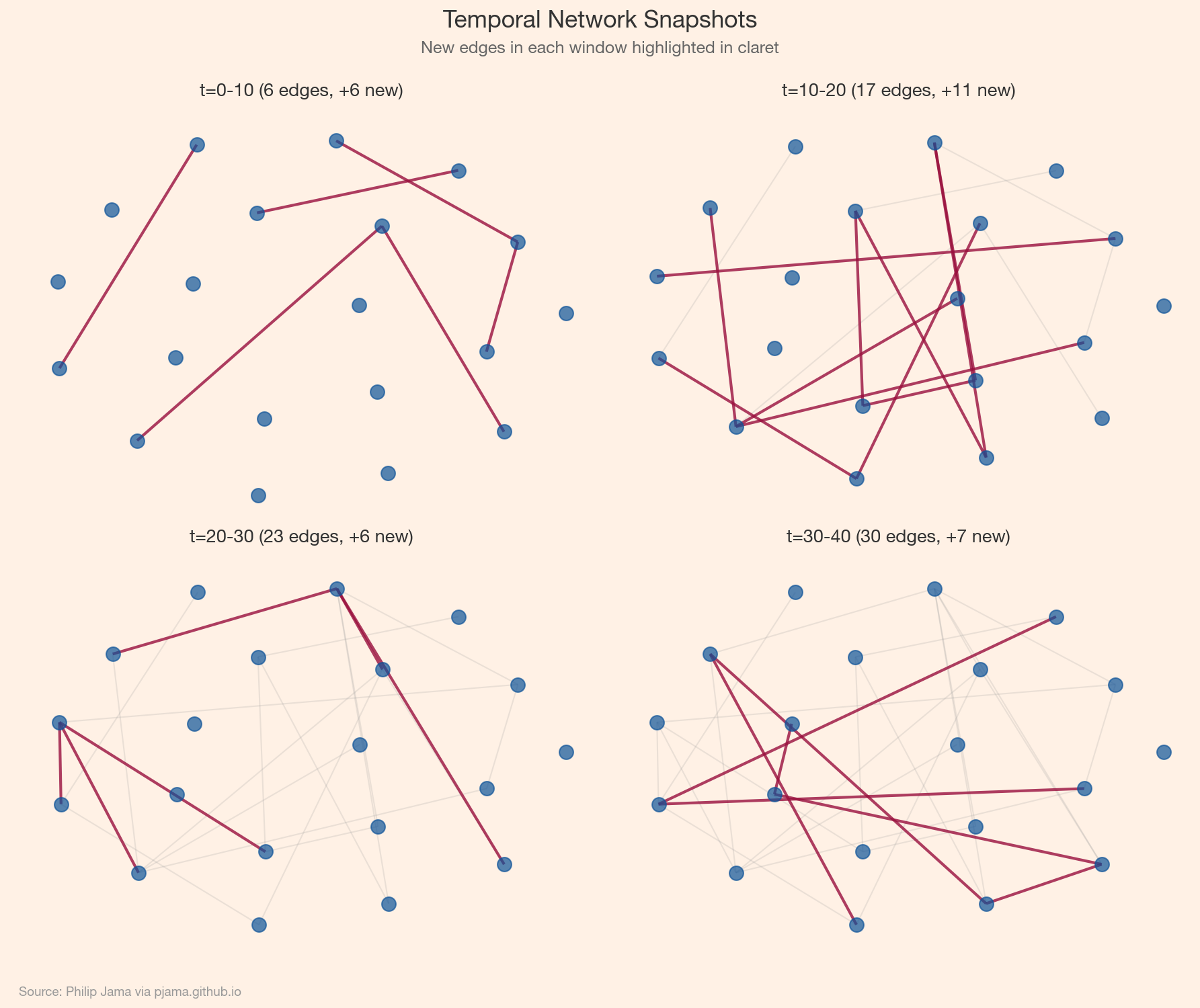
Standard random walks ignore time: a walker can traverse an edge from 2020 and then one from 2015. Temporal random walks enforce chronological ordering: each step must follow an edge with a timestamp later than the previous step. This produces time-respecting paths that capture causal influence and temporal reachability. Temporal node2vec extends this to learn time-aware node embeddings.
TGAT (Temporal Graph Attention) applies attention over time-stamped neighbors, weighting recent interactions more heavily using time-encoding functions. TGN (Temporal Graph Networks) adds a memory module: each node maintains a state vector updated after each interaction, allowing the model to remember long-term patterns. Both process events sequentially and produce dynamic node embeddings suitable for link prediction and anomaly detection.
Consider an email or messaging dataset: each message is a timestamped directed edge. A temporal graph model can predict who will message whom next, detect emerging communities, or flag anomalous communication bursts. The evolution of network metrics over time: density, clustering, reciprocity. That evolution tells a story that no single snapshot captures.
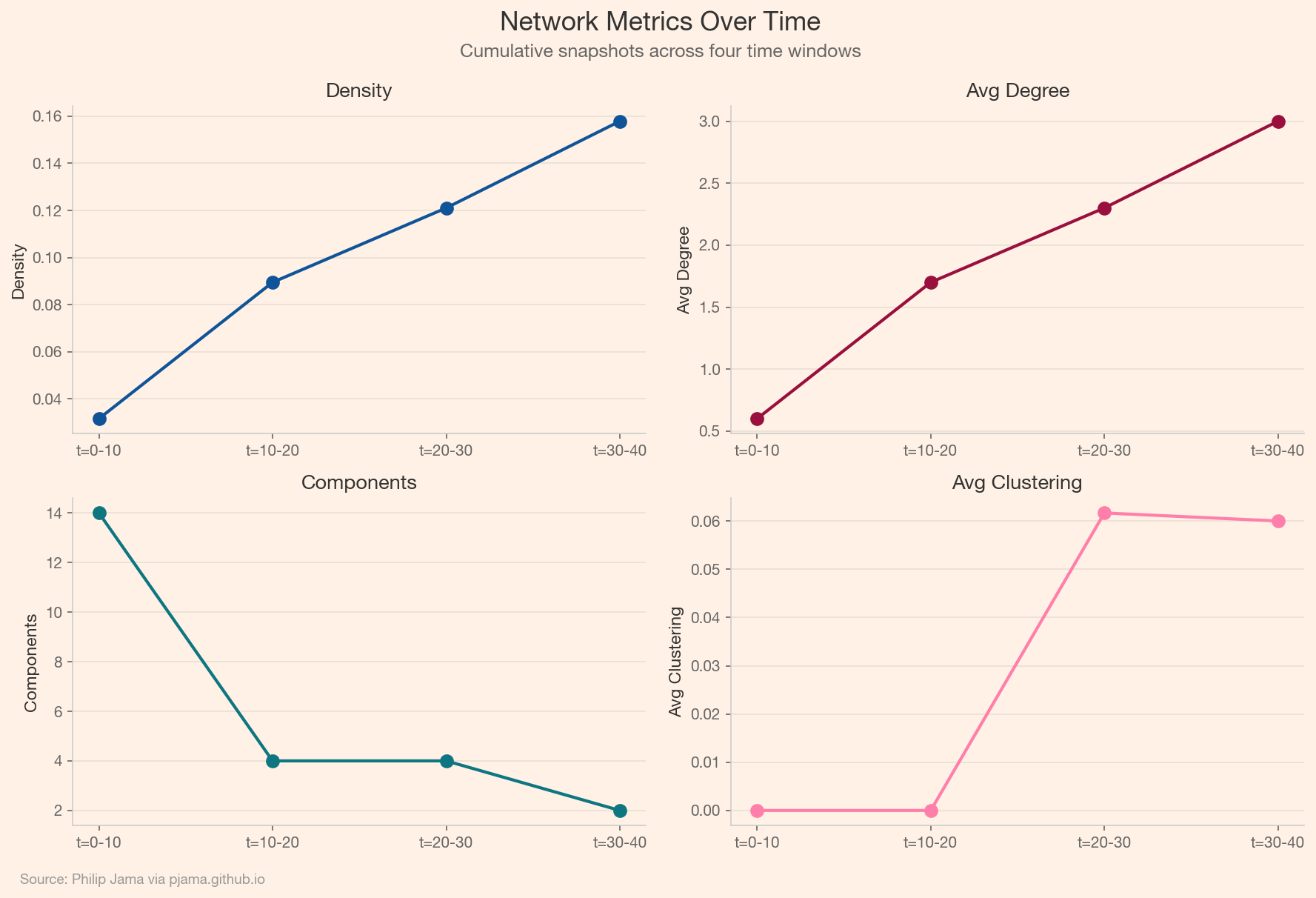
Consider a B2B services company where account managers, support engineers, and executives communicate with client contacts via email, calls, and meetings. Each interaction is a timestamped edge between an internal employee and an external contact, carrying metadata: channel, duration, topic tags, sentiment. The resulting temporal bipartite graph encodes the full relationship history between the company and its customers.
A healthy account has regular, multi-threaded communication: the account manager checks in, support resolves tickets, an executive joins quarterly reviews. When those threads start thinning -- fewer touchpoints, longer gaps between interactions, conversations narrowing to a single channel -- the temporal pattern signals risk before any explicit complaint arrives.
Link prediction on this graph asks: given the communication history up to time t, which edges are likely to occur at t+1? A TGN trained on historical account data learns the cadence of healthy relationships. When the model predicts a touchpoint that fails to materialize -- a quarterly review that doesn't happen, a support thread that goes unanswered -- the gap between prediction and reality becomes a churn signal. The account team can intervene before the silence becomes a cancellation.
The same graph supports anomaly detection from the opposite direction. Instead of predicting missing edges, flag edges that shouldn't be there -- or that deviate sharply from the learned pattern. A sudden spike in support tickets from a previously stable account, a burst of escalation emails bypassing the usual contacts, or a dormant executive relationship that abruptly reactivates: these are temporal anomalies that carry operational meaning.
The temporal dimension is essential here. A single support ticket is routine. Five support tickets in a week from an account that averages one per month is an anomaly that only surfaces when the model understands the baseline rhythm. Static graph analysis would count edges; temporal analysis understands tempo.
The practical value comes from connecting graph signals to business workflows. Link prediction scores can feed a CRM dashboard that ranks accounts by engagement risk. Anomaly scores can trigger alerts routed to the relevant account manager. Temporal community detection -- identifying clusters of client contacts who interact with different internal teams -- can surface accounts where communication is siloed and a single point of failure exists. The graph does not make the decision, but it surfaces the pattern that would otherwise remain buried in thousands of individual interactions.
Temporal networks model how graphs change but not why. Causal and Bayesian networks add directionality and interventional reasoning.
If you're exploring related work and need hands-on help, I'm open to consulting and advisory. Get in touch›