Articles /Network Graph Analysis /Part 9
Causal and Bayesian Graph Networks
DAGs, interventions, and counterfactual reasoning on graphs
Causal InferenceBayesian NetworksDAGDo-CalculusCounterfactualsPythonProbabilistic Graphical Models
Articles /Network Graph Analysis /Part 9
Causal InferenceBayesian NetworksDAGDo-CalculusCounterfactualsPythonProbabilistic Graphical Models
Temporal models capture when connections change but not why. Causal and Bayesian graph networks use directed acyclic graphs to encode cause-and-effect, turning the graph from a descriptive tool into a reasoning engine. Where Part 8 (Temporal Graph Networks) modeled evolving structure, this final article asks what that structure means: which edges represent genuine causal influence, and what would happen if we intervened on a node.
The framework comes from Judea Pearl’s causal inference tradition. We start with the DAG as a causal model, develop the do-calculus for reasoning about interventions, add Bayesian networks for probabilistic inference, and close with counterfactual reasoning -- the strongest claim a graph can support.
In a causal DAG, nodes represent variables and directed edges represent direct causal effects. The absence of an edge is an assertion: it claims one variable does not directly cause another. This graphical language, formalized by Judea Pearl, encodes three fundamental structures.
A confounder is a common cause of two variables. If socioeconomic status causes both education level and health outcome, the edge from status to each creates a spurious association between education and health. A mediator lies on the causal path: a drug affects gene expression, which affects tumor size. The drug’s effect on the tumor is mediated through gene expression. A collider is a common effect of two variables. If both talent and attractiveness independently increase the chance of becoming a celebrity, conditioning on celebrity status creates a spurious negative association between talent and attractiveness -- the phenomenon known as Berkson’s paradox.
Each structure has distinct implications for statistical analysis. Confounders must be adjusted for, mediators define indirect effects, and colliders must not be conditioned on. The DAG makes these relationships explicit and testable.
These three structures determine what a DAG can tell you about conditional independence -- a property called d-separation. By tracing paths through the graph and checking whether intermediate nodes are conditioned on, you can read off which variables are independent given what you observe. Conditioning on a confounder blocks the spurious path. Conditioning on a collider opens one. This is why the observational causal inference article emphasized drawing the DAG before choosing which variables to adjust for: the graph’s structure dictates what to condition on, and getting it wrong introduces bias rather than removing it.
D-separation also gives the DAG testable implications. Every independence it predicts can be checked against data. If the data show an association where the DAG predicts independence, the DAG is wrong -- a property that structure learning algorithms exploit to discover causal graphs from data.
Observational data tells us $P(Y \mid X = x)$ -- the probability of $Y$ given that we observe $X$ taking value $x$. But the causal question is $P(Y \mid \text{do}(X = x))$ -- the probability of $Y$ if we set $X$ to $x$ by intervention. Pearl’s do-operator formalizes this distinction through graphical surgery: to compute the effect of $\text{do}(X = x)$, delete all incoming edges to $X$ in the DAG (since the intervention overrides whatever would have caused $X$) and compute the resulting distribution.
The key question is when we can compute this interventional distribution from observational data alone. The backdoor criterion provides the most common answer. In a causal DAG, some paths between treatment and outcome carry the causal effect (they flow forward through direct or mediated connections). Others are confounding paths: they flow from a common cause into both treatment and outcome, creating a spurious association. If we can find a set of variables $Z$ that blocks all confounding paths without opening new ones (by avoiding conditioning on colliders), the causal effect reduces to a conditional probability weighted over $Z$:
$$P(Y \mid \text{do}(X = x)) = \sum_z P(Y \mid X = x, Z = z) \; P(Z = z)$$
The front-door criterion covers a different situation: when confounders between treatment and outcome are unmeasured but the entire causal effect flows through an observed mediator. If we can estimate the effect of treatment on the mediator and the effect of the mediator on the outcome separately, we can chain them to recover the total causal effect -- even without measuring confounders directly. The following visualization shows the graphical surgery that distinguishes observation from intervention.
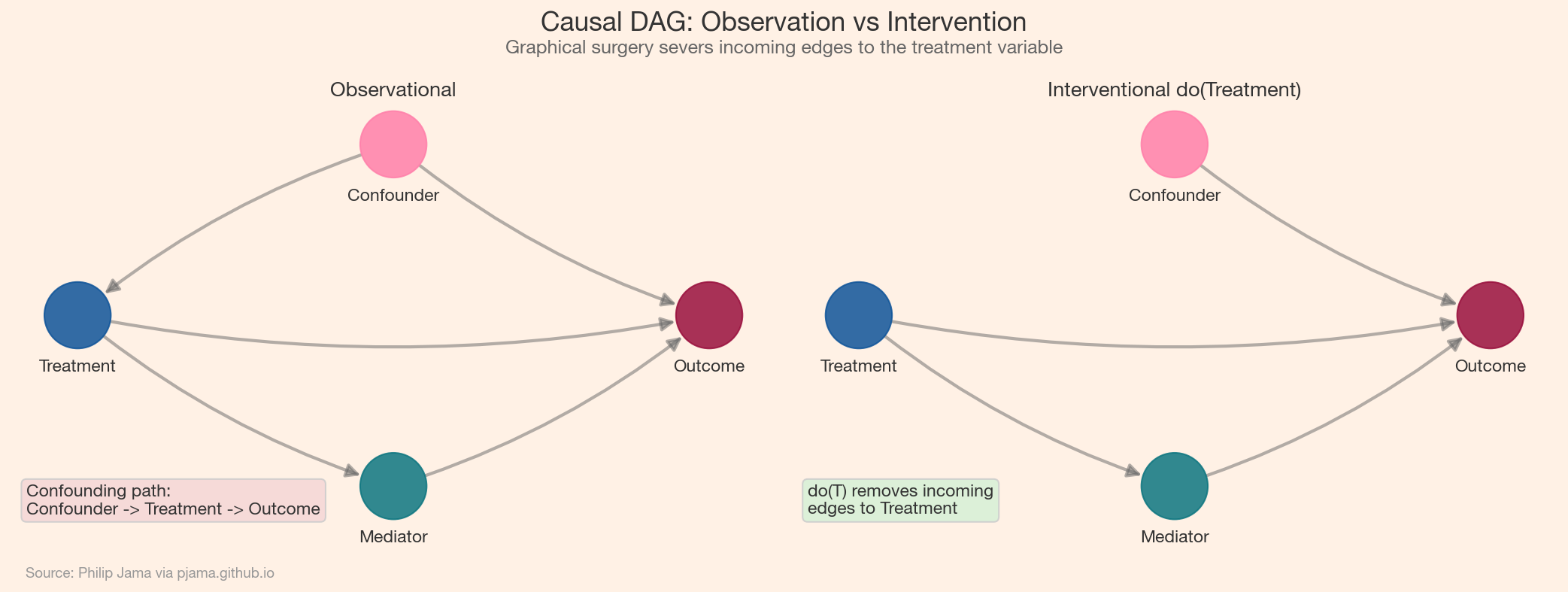
Randomized experiments implement the do-operator physically. When we randomly assign subjects to treatment or control, we sever the connection between the treatment variable and any confounders -- exactly the graphical surgery that do(X = x) performs on the DAG. This is why randomization eliminates confounding bias without requiring us to identify or measure confounders.
The Online Experiments with a Bayesian Lens article analyzed A/B tests from this perspective: random assignment guarantees that the backdoor criterion is satisfied with Z = ∅, so the observed difference in outcomes is the causal effect. When randomization is impossible -- in observational studies, natural experiments, or retrospective analyses -- the do-calculus provides the machinery for determining whether and how the causal effect can still be recovered.
A Bayesian network pairs a DAG with a set of conditional probability distributions (CPDs): each node has a distribution conditioned on its parents. Together, these local specifications define a joint distribution over all variables through the factorization property: $P(X_1, \ldots, X_n) = \prod_i P(X_i \mid \text{Parents}(X_i))$. The DAG structure determines which conditional independencies hold, and the factorization exploits them to represent a potentially huge joint distribution compactly.
The classic textbook example uses three variables: Rain, Sprinkler, and Grass Wet. Rain influences both the sprinkler (negatively -- you turn off the sprinkler when it rains) and whether the grass is wet. The sprinkler also affects whether the grass is wet. Given the observation that the grass is wet, we want the posterior probability that it rained -- a computation that requires combining the prior on rain with the likelihoods through each causal pathway.
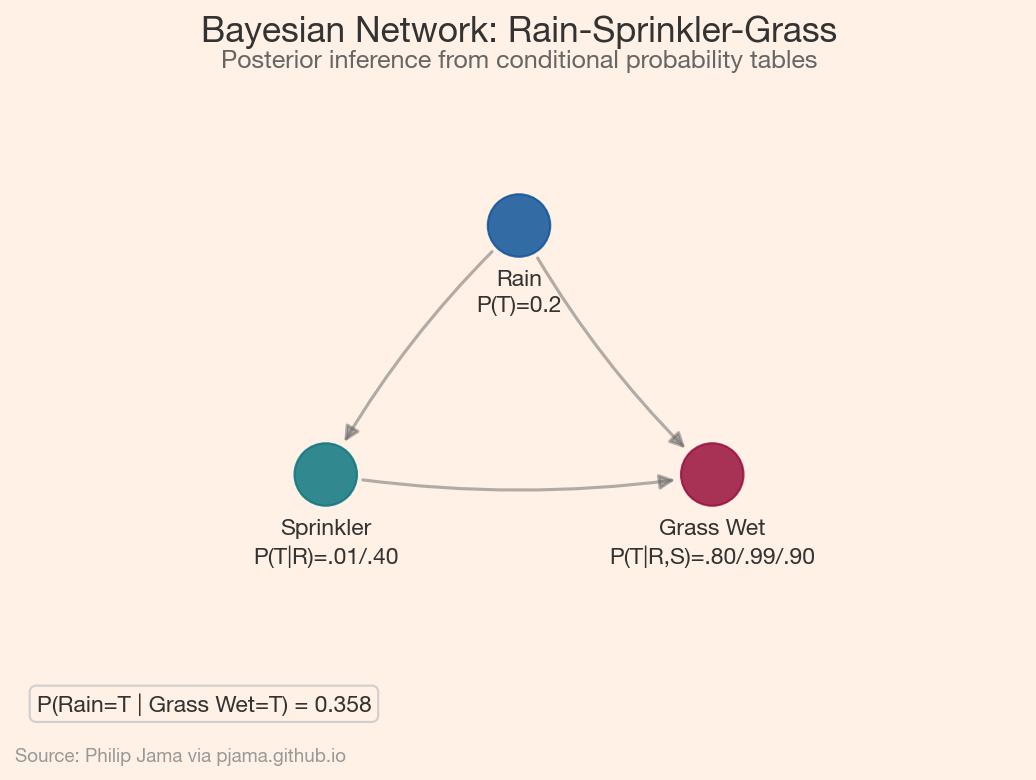

Inference in Bayesian networks, computing posterior probabilities given observed evidence, exploits the graph structure for efficiency. Exact methods like variable elimination marginalize out variables one at a time, choosing an elimination order guided by the DAG’s topology. When exact inference becomes intractable (large graphs, continuous variables), approximate methods like MCMC sampling or variational inference step in. The DAG determines which approach is feasible: sparse graphs with few connections per node admit exact solutions, while dense networks require approximation.
In practice, the causal DAG is rarely known with certainty. Structure learning algorithms attempt to recover the graph from data. The fundamental limitation is that observational data alone cannot distinguish between DAGs that encode the same set of conditional independencies. Such DAGs form a Markov equivalence class: they share the same edges but disagree on the direction of some arrows, and no observational test can tell them apart. Structure learning can narrow the possibilities to this class but cannot pinpoint the unique causal DAG without additional assumptions or experimental data.
Two families of algorithms dominate. Constraint-based methods like the PC algorithm test conditional independencies in the data, remove edges between independent variables, and orient the remaining edges using structural patterns (e.g., if two non-adjacent nodes both point to a third, that collision pattern reveals edge direction). Score-based methods like greedy equivalence search (GES) search over graph structures to optimize a fit-complexity tradeoff, typically using the Bayesian Information Criterion. Hybrid approaches combine both strategies.
In Python, pgmpy provides both the PC algorithm and Bayesian network inference in a unified API. CausalNex adds continuous optimization methods and integrates domain knowledge through edge whitelisting and blacklisting. DoWhy focuses on causal effect estimation, wrapping identification and refutation steps around a user-specified or learned DAG. In practice, purely data-driven learning is a starting point: domain experts fix edges that are known (temporal ordering, established mechanisms) and let the algorithm fill in uncertain relationships.
Consider a microservices architecture where Service A (API gateway) calls Service B (authentication) and Service C (product catalog), which in turn calls Service D (database layer). Distributed tracing produces a directed call graph with latency measurements at each edge. When end-to-end latency spikes, which service is the root cause?
Framing this as a causal problem: the call graph is a causal DAG, where each node’s latency is a function of its own processing time plus the latencies of its downstream dependencies. A spike in Service D’s latency causally propagates to C and then to A. But the system also has confounders -- shared infrastructure like CPU and network bandwidth that affect multiple services simultaneously. The raw correlation between Service B’s latency and user-facing latency might be driven entirely by a shared load balancer, not by any causal path through B.
Pearl’s causal ladder distinguishes three levels of reasoning, each strictly more powerful than the last. Association (level 1) asks "what is": given that Service A is slow, how likely is Service D to be slow? This is standard observational correlation. Intervention (level 2) asks "what if we do": if we force Service D’s latency to its baseline (do(D = normal)), does user-facing latency recover? This requires the do-calculus machinery developed above. Counterfactual (level 3) asks "what if things had been different": given that the outage did occur, would it have occurred if Service D had not exceeded its memory limit?
Counterfactual reasoning follows a three-step procedure. First, abduction: use the observed evidence (all measured latencies during the incident) to infer the values of unobserved variables (network congestion, background load). Second, action: modify the DAG to reflect the hypothetical intervention (set Service D’s memory usage to normal). Third, prediction: propagate forward through the modified DAG to compute the counterfactual outcome. The result is an attribution score: the probability that the outage would not have occurred in the counterfactual world.
D-separation provides the first filter. If Service B and the user-facing latency are d-separated given the gateway’s processing time, then B cannot be the root cause regardless of its observed latency -- any correlation is explained by the conditioning set. This narrows the candidate set without any statistical estimation, using only the graph structure.
For the remaining candidates, the do-calculus formalizes the root cause question. The interventional query P(user_latency > threshold | do(C = normal)) asks: if we could reset Service C’s latency, how much would user-facing latency improve? If the backdoor criterion is satisfied (by conditioning on shared infrastructure metrics), this quantity can be estimated from the observational traces without actually intervening on the running system.
The counterfactual step completes the analysis. After the incident, the post-mortem asks: "Would the outage have occurred if Service D had not exceeded its memory limit?" The three-step procedure -- abduct the hidden variables from the incident data, impose do(D_memory = normal), predict the resulting latencies -- produces a probability. If it is low, Service D’s memory spike was the root cause. If it is high, the outage would have happened anyway, and the true cause lies elsewhere in the graph.
This layered approach -- association for monitoring, intervention for diagnosis, counterfactual for attribution -- mirrors how incident response teams actually reason. The causal graph formalizes each step and makes the assumptions explicit. When those assumptions are encoded in a DAG, they become auditable: anyone can inspect the graph, check the d-separation claims, and challenge the causal model.
This series has moved through progressively stronger claims about what graphs can tell us. Part 1 (Foundations) defined the vocabulary: nodes, edges, paths, components. Subsequent articles measured structure (centrality, communities), extracted it from text (knowledge graphs), learned representations of it (GNNs), and tracked how it changes (temporal networks). Each step added analytical power while staying within the realm of association -- describing patterns in data.
Causal and Bayesian graph networks cross that boundary. The DAG is no longer a summary of observed relationships but a claim about the data-generating process. D-separation produces testable predictions. The do-calculus turns observational data into interventional conclusions. Counterfactuals attribute specific outcomes to specific causes. The shift from association to causation is what separates description from understanding, and graphs are the language in which that shift is expressed.
The causal reasoning developed here connects directly to the Decision Science series: Bayesian A/B testing is intervention with randomization, observational causal inference is intervention without it, and the DAG framework unifies both.
If you're exploring related work and need hands-on help, I'm open to consulting and advisory. Get in touch›