Articles /Network Graph Analysis /Part 1
Foundations
Graph representations, key metrics, and visualization with NetworkX
Graph TheoryPythonNetworkXTutorial
Articles /Network Graph Analysis /Part 1
Graph TheoryPythonNetworkXTutorial
Graphs are everywhere: social networks, transportation routes, biological pathways, the internet itself. A graph is simply a set of nodes connected by edges, yet this minimal structure encodes rich relational information that matrices and tables miss. This article lays the groundwork for a series on network graph analysis: starting with the representations, metrics, and visualizations you need to reason about any network.
The series draws on three portfolio projects that use graph techniques: Calendar Graph project ↗ (organizational network analysis), Graphception project ↗ (concept extraction and associative networks), and Books project ↗ (LLM-generated outlines as tree graphs).
A graph G = (V, E) consists of a vertex set V and an edge set E. Edges can be directed or undirected, weighted or unweighted. This abstraction lets us model friendships, citations, dependencies, and countless other relationships in a single framework.
How you store a graph matters. An adjacency matrix is an n×n array where entry (i,j) indicates an edge between nodes i and j: great for dense graphs and linear algebra. An edge list is a simple collection of (u, v) pairs: compact and easy to stream. An adjacency list maps each node to its neighbors: ideal for sparse graphs and traversal algorithms. NetworkX abstracts over these, but understanding the tradeoffs guides performance decisions.
A handful of metrics capture a graph’s structure:
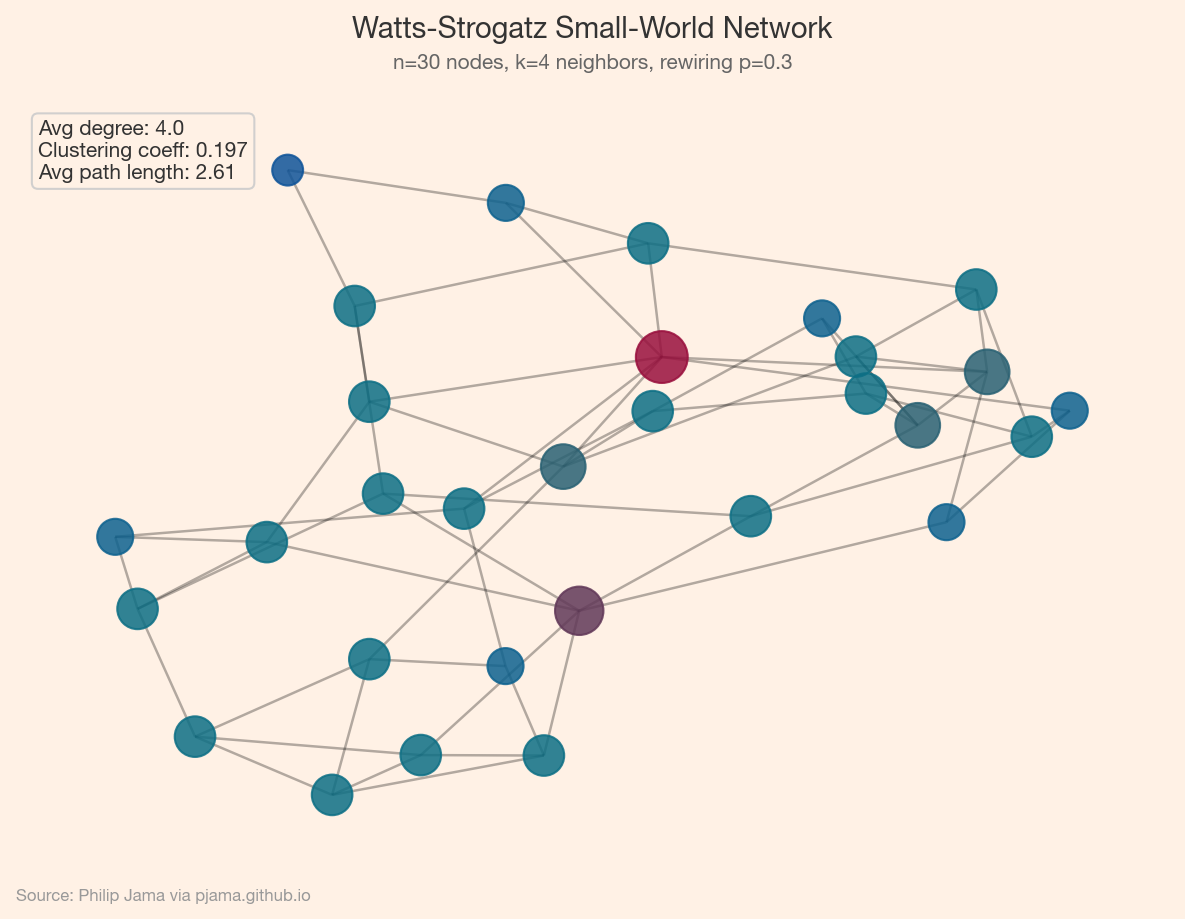
Layout algorithms turn abstract graphs into spatial pictures:
No layout is objectively best: each reveals different aspects of the same graph.
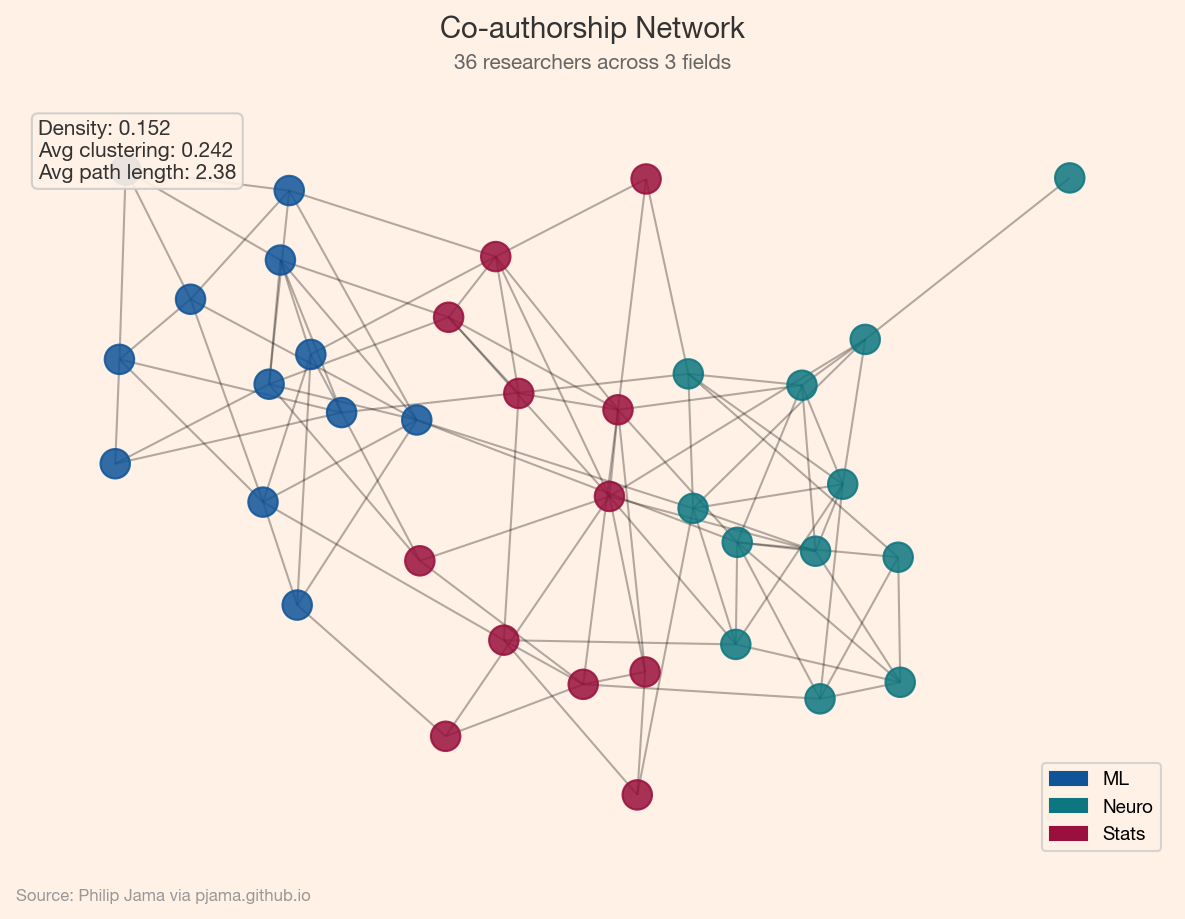
With these foundations in place, the series branches into three directions:
Branch A: Social & Organizational Networks
Branch B: Knowledge Graphs
Branch C: Graph Deep Learning
Each branch builds on the representations and metrics introduced here.
Degree distributions and clustering coefficients describe a graph's texture -- but the most useful structure is often hidden in densely connected subgroups.
If you're exploring related work and need hands-on help, I'm open to consulting and advisory. Get in touch›